Systematic review on the known variants and genes associated with orofacial clefts in Africa
Thirona Naicker, Azeez Alade, Waheed Awotoye, Helen Malherbe, Colleen Aldous, Azeez Butali
Corresponding author: Thirona Naicker, Genetics, Genetics, Department of Paediatrics, University of KwaZulu-Natal and Inkosi Albert Luthuli Central Hospital, Durban, South Africa 
Received: 25 Sep 2021 - Accepted: 01 Aug 2022 - Published: 30 Aug 2022
Domain: Genetics ,Pediatric Dentistry,Health Research
Keywords: Systematic review, orofacial clefts, gene variants, Africa
©Thirona Naicker et al. PAMJ Clinical Medicine (ISSN: 2707-2797). This is an Open Access article distributed under the terms of the Creative Commons Attribution International 4.0 License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Cite this article: Thirona Naicker et al. Systematic review on the known variants and genes associated with orofacial clefts in Africa. PAMJ Clinical Medicine. 2022;9:46. [doi: 10.11604/pamj-cm.2022.9.46.31756]
Available online at: https://www.clinical-medicine.panafrican-med-journal.com//content/article/9/46/full
Systematic review on the known variants and genes associated with orofacial clefts in Africa
![]() Thirona Naicker1,2,
Thirona Naicker1,2, ![]() Azeez Alade3, Waheed Awotoye4, Helen Malherbe5,
Azeez Alade3, Waheed Awotoye4, Helen Malherbe5, ![]() Colleen Aldous6, Azeez Butali2,7
Colleen Aldous6, Azeez Butali2,7
&Corresponding author
A systematic review of the literature was conducted to assess the known variants and genes associated with orofacial clefts (OFC) in Africa. This was performed via an electronic search, using a set of predefined search terms, in seven databases (PubMed, EBSCOhost, ScienceDirect, Web of Science, Google Scholar, OpenGrey and WorldCat) from 2000-2020. Full text articles concerning the genetic aetiology of OFC in Africa were included. Articles were filtered using the PICO eligibility criteria and data extraction followed the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines. The primary search yielded 489 citations, of which 16 studies were finally analyzed. We identified 61 genetic variants across the 16 studies, and 35 were novel variants. Interferon Regulatory Factor 6 (IRF6) was the most common gene investigated. In most studies, the genetic testing was not performed in Africa. There is a wealth of information to be learnt from interrogating the African genome as we try to understand the role of variants in health and diseases. The protocol for this systematic review is registered in the international prospective register of systematic reviews (PROSPERO CRD42020199361).
Africa is the second largest and the second most populous continent in the world, after Asia [1], with the population of sub-Saharan Africa projected to triple by the year 2100 [2,3]. Today, an estimated 1.34 billion [4] people live on the African continent and due to the socio-economic status of the 54 countries in Africa, many are classified as low-middle income countries (LMIC). As such, there are budgetary constraints for many essential services, including health and education. Historically, African health care budgets focused on medical conditions that resulted from a high burden of disease. Prevention of communicable, infectious diseases such as Human Immunodeficiency Virus (HIV) and tuberculosis (TB) is an important strategy for primary health care in developing countries [5]. Hence, genetic testing and genomic literacy for non-communicable diseases (NCDs) are poorly developed in Africa. Congenital disorders (CDs), the first NCDs experienced in life, are defined as any potential pathological condition present at birth, whether caused by environmental, genetic or unknown factors [6]. Congenital disorders account for a large proportion of child and infant morbidity and mortality. According to the 2006 March of Dimes global report on birth defects, more than 7.9 million children are born every year with severe genetic conditions or malformations. Ninety percent of these children are born in LMIC [7]. Updated global estimates (2010-2014) indicate that CDs annually account for over 400,000 foetal deaths, 2.5 million under-5 deaths and a further 2 million children surviving with significant, lifelong disability [8,9].
One of the most obvious CDs present across all populations are orofacial clefts (OFC), which occurs with an estimated birth prevalence rate of 0.5 per 1000 live births or 1:2000 children in Africa [10]. Although research in Africa is promising, limited work has been conducted on the epidemiology and genetic aetiology of OFC in the continent. Regarding OFC research in LMICs, a recent scoping review looked at six hundred and twenty publications over the 10-year period. More than half of the articles were reported from Asia (57%), then from Americas (22.8%), Africa (15.4%) and the rest from cross-regional, Europe and Oceania (4.9%) [11]. Individuals of African ancestry are significantly underrepresented in clinical research and initiatives such as the Human Heredity and Health in Africa (H3Africa) and other projects facilitating the discovery of novel genomic loci in Africans [12]. Genetic causes of OFC in Africa are not fully understood. Various unrelated studies carried out across the continent show the diversity of the African genome in patients with cleft lip with or without cleft palate (CL±P). Several novel variants have been found, indicating that the African genome needs to be further explored for the benefit of those affected by OFC on the continent and to provide further insight into known genetic causes. The objective of this systematic review was to understand and document the research studies that have been undertaken in the field of OFC utilizing genetic testing to determine the known variants and genes associated with OFC in Africa in the last 20 years, specifically 2000 - 2020.
Protocol and registration: the protocol for this review i.e., the inclusion criteria and methodology, was designed prior to data collection and registered in the international prospective register of systematic reviews (PROSPERO) [13], with registration number: CRD42020199361. Reporting followed the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines [14].
Eligibility criteria: this review included peer-reviewed papers on humans, available in English from 2000 until 2020. The following PICO eligibility criteria were utilized: (P) population: patients with OFC in Africa; (I) intervention: analysis of genetic variants and genes; (C) comparison: related (case triads) and unrelated controls; (O) outcome: analysis of the variant´s predicted pathogenicity, i.e. CADD scores and other variant prediction tools.
Information sources: an electronic search using PubMed, EBSCOhost (including MEDLINE and CINAHL), ScienceDirect, Web of Science and Google Scholar databases was conducted. In addition, grey literature was searched through OpenGrey and WorldCat.
Search strategy: the Boolean search strategy consisted of the following MESH terms: cleft OR Clefting OR clefts OR "cleft*" AND Lip OR Palate OR Orofacial OR Craniofacial OR facial; syndromic or nonsyndromic; genome OR genotype OR genotyping OR "genotyp*" OR genetics OR genetic OR (genetic and analysis) OR (genetics and studies) OR (genome and wide and association) and (study or studies)) OR GWAS OR genes or gene OR hereditary OR inherited OR variants OR sequencing OR mutation OR SNPs OR “Single nucleotide polymorphism” Africa
Study selection: publications from all database searches were imported into the Endnote X9 reference manager [15] where duplicate records were removed. Two authors (TN and AAA) independently reviewed the titles, keywords, and abstracts of all selected articles to assess whether the studies were eligible for inclusion in the review. A Cohen´s Kappa score of 0.86 was achieved when the title and abstracts were screened. Full text of the selected articles was retrieved (where available) and reviewed to confirm eligibility by the same two authors, resulting in, a Kappa score of 0.76. This showed substantial agreement, and all disagreements were resolved by discussion between the two reviewing authors (TN and AAA) and a third senior reviewer (AB).
Data collection process and items: data was extracted into a data abstraction form developed in Microsoft Excel. Information collected from each study included: year of publication, first author´s name, title, study design, location of the study, number of participants and controls, phenotypic data including cleft type and whether syndromic or non-syndromic, genomic analyses/test, laboratory location, variant detected and a functional variant assessment (if included in the study). The Combined Annotation-Dependent Depletion (CADD) scores and American College of Medical Genetics (ACMG) classification were computed for variants that did not have them.
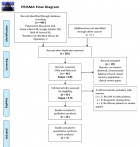
The electronic search of the databases resulted in a total of 489 citations. Twenty-four were retrieved from PubMed, 124 from Ebscohost, 84 from Science Direct, 90 from Web of Science and 139 from Google Scholar. Grey literature searches provided 2 citations from OpenGrey, 12 from World Catalogue and 14 from World Catalogue Thesis. No additional records were found through other sources. After elimination of duplicate records, 395 remaining citations were screened based on titles, abstracts, and full text review. Sixteen articles met all the inclusion criteria and were finally analyzed in this systematic review. The PRISMA flow diagram is shown in Figure 1. The 16 included studies were published between 2005 and 2020 with 15 (94%) of these studies published in the last nine years, since 2010 (Annex 1). Studies were classified as case-control or family-based studies, with some studies having both. There was only one African genome wide association study (GWAS) published in 2019 [16]. Patients from Nigeria, Ghana and Ethiopia were featured in nine studies. Eight (50%) of the studies included non-syndromic OFC patients only, while seven included syndromic patients only and one study included both. Most studies (11/16) utilized Sanger sequencing of candidate gene(s) as the method to identify the variants. One study conducted whole genome sequencing using the expanded Illumina MEGA v2 with imputation, and another study conducted whole exome sequencing. Family-based association tests, single variant association tests of independent samples, meta-analysis and segregation analysis were the common analyses methods employed. There were 61 variants described across the 16 studies (Annex 1). A total of 24 of these were classified as missense variants and 35 were novel. IRF6 was investigated in five of the studies, and hence was the most common gene studied. Other common candidate genes researched in African papers were PAX7, 8q24, ARGHAP 29, MSX1, and VAX1.
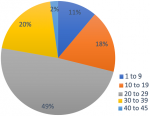
In 14 of the 16 studies (88%), the genetic testing was performed in laboratories outside of Africa. Nine of these studies (64%) were performed in Iowa, USA. Genetic testing in cleft patients were only conducted in North African countries, Tunisia, and Morocco in in 2005 and 2014; respectively. Seven studies included functional assessments of the variants found. The CADD scores, which categorise genetic variants (single nucleotide, short insertions, and deletions) throughout the human genome, ranged from 1.12 to 40 with many scores being above 20. The top ranked variants show CADD scores of greater than 10, 20 or 30 which means that the variant falls in the top 10%, 1% or 0.1% of the most deleterious mutations in the human genome, respectively. Of the African variants with calculable CADD scores; 22/45 (49%) had scores of greater than 20, nine variants (20%) had CADD scores greater than 30, and one variant had a score of 40 (Annex 1). These scores place the variants in the top 1%; 0.1% and 0.01% of deleterious mutations respectively; making them highly significant (Figure 2). According to the American College of Medical Genetics (ACMG) variant classification, 23/61 (38%) variants are pathogenic or likely pathogenic and may likely contribute to the cleft phenotype. A further 18/61 (30%) variants are variants of unknown significance (VUS) and there will be a need to reclassify them in a year or two once more information becomes available. This review explores what is known about OFC variants and genes in Africa. Over a 20-year period, 16 studies conducted genetic testing on OFC patients.
Nigeria, Ghana, and Ethiopia form part of the African Craniofacial Anomalies Research Network (AfriCRAN) and featured in 9/16 (56%) published African studies. This network, which was conceptualised in 2012, represents a group of craniofacial researchers in Africa who work with international collaborators to investigate the genetic and environmental aetiology of OFC in their countries [17]. African populations display vast genetic diversity [18] and we found that 35/61(57,3%) were novel variants, showing that the African genome has yet to be completely interrogated. These variants were novel at the time of publication, however as more sequencing data becomes available from Africa and the studies evolve, we will discover that those variants are no longer novel. The importance of identifying novel variants of OFC in Africa is directly related to the care and prevention of those affected by OFC. By identifying the African OFC variants, we can counsel patients appropriately regarding the mode of inheritance and possibly prevent recurrence in future generations through prenatal testing.
Whilst there was only one African cleft GWAS study conducted to date [16], which included 1019 patients and 2159 controls; this study only interrogated patients from Nigeria, Ghana and Ethiopia and therefore did not represent all African countries. There is a need for more omics studies to be conducted to better understand the genetic and genomic causes of clefts from other countries in Africa. Only 12.5% of studies (2/16) utilised Sanger sequencing available in Africa. While encouraging, this represents a small proportion and demonstrates the limited capacity for genomic testing in Africa. While costs of genetic testing are decreasing globally, the cost of sequencing is considerable and grant funders and governments need to consider investment to train and build capacity in Africa. Despite the limitations, African researchers have succeeded in obtaining diagnoses for their patients, due to collaboration with international colleagues and laboratories. This shows that skilled human resources are present in Africa but the supporting infrastructure (and political commitment to implement these technologies) is lacking.
African governments require a mindset change around the funding of genetic research. A recent report on The Economic Impact and Functional Applications of Human Genetics and Genomics [19] showed that the field of human genomics and genetics supported 850,000 jobs and generated $265 billion across the United States (US) economy in 2019. The potential socioeconomic benefits of implementing these technologies across Africa need further investigation. One can only imagine how much can be generated and the number of jobs that can be created if the right investments are made in Africa. Until this is achieved, Africa will continue to remain a consumer continent, rather than investing in research and aiming to become a discovery continent. This is best illustrated by the current COVID-19 pandemic where African countries needed to raise funds to buy the internationally manufactured vaccine for their citizens as they were unable to produce the required doses in-country.
Limitations: this systematic review provides an overview of the OFC variants found in Africa, limited by different testing methods and study types across the 16 studies. In addition, functional assessments or CADD scores of the variants were not computed for all studies. Although this review intended to find all the available research on the genetic aetiology of OFC in Africa, it is possible that some studies were not detected.
This study highlights that more local genetics and genomics studies involving Africans need to be conducted. These studies will contribute substantially to our understanding of health and diseases.
Funding: the authors disclosed receipt of the following financial support for the research. This work was supported by funding from Smile Train and the National Institute of Health/National Institute of Dental and Craniofacial Research (R00 DE022378 and R01 DE28300).
What is known about this topic
- We know that causative variants and genes discovered in European and Asian populations cannot be extrapolated into African populations;
- There are novel variants to be found in African populations;
- The African genome is an untapped resource and there is still much to understand about human origin, diversity, and disease susceptibility.
What this study adds
- This review collates all the variants from African orofacial cleft studies over a 20-year period.
- We utilise bioinformatic scoring tools like CADD and the ACMG classification to analyse and determine if the variants can result in pathology.
- It brings to attention that African governments need to invest in genomic research and become a continent that makes new discoveries.
The authors declare no competing interests.
TN, AAA, AB contributed to the study design, acquisition of data, analysis, and interpretation of data, wrote and revised the final manuscript. WA contributed to analysis, interpretation of data and the revision of the final manuscript. CA contributed to the study design and revision of the final manuscript. HM contributed to the revision of the final manuscript.
Figure 1: PRISMA flow diagram
Figure 2: CADD score distribution
Annex 1: showing studies 1-16 and variant details
- UN. World Population Prospects 2019. 2019.
- Ezeh A, Kissling F, Singer P. Why sub-Saharan Africa might exceed its projected population size by 2100. Lancet. 2020 Oct 17;396(10258):1131-3. PubMed | Google Scholar
- Vollset SE, Goren E, Yuan CW, Cao J, Smith AE, Hsiao T et al. Fertility, mortality, migration, and population scenarios for 195 countries and territories from 2017 to 2100: a forecasting analysis for the Global Burden of Disease Study. Lancet. 2020 Oct 17;396(10258):1285-306. PubMed | Google Scholar
- Worldometer. Africa population. 2021. Accessed Sept 25, 2021.
- Bradshaw D, Groenewald P, Laubscher R, Nannan N, Nojilana B, Norman R et al. Initial burden of disease estimates for South Africa, 2000. S Afr Med J. 2003 Sep;93(9):682-8. PubMed | Google Scholar
- Joint W, Organization WH. Management of birth defects and haemoglobin disorders: report of a joint WHO-March of Dimes meeting. Geneva, Switzerland, 17-19 May 2006. Google Scholar
- Christianson A, Howson CP, Modell B. March of Dimes: global report on birth defects, the hidden toll of dying and disabled children. March of Dimes: global report on birth defects, the hidden toll of dying and disabled children. 2005. Google Scholar
- Modell B, Darlison M, Moorthie S, Blencowe H, Petrou M, Lawn J. Epidemiological methods in community genetics and the Modell Global Database of Congenital Disorders (MGDb). UCL. 2016. Google Scholar
- Saib MZ, Dhada BL, Aldous C, Malherbe HL. Observed birth prevalence of congenital anomalies among live births at a regional facility in KwaZulu Natal Province, South Africa. PLoS One. 2021;16(8):e0255456. PubMed | Google Scholar
- Butali A, Adeyemo WL, Mossey PA, Olasoji HO, Onah, II, Adebola A et al. Prevalence of orofacial clefts in Nigeria. Cleft Palate Craniofac J. 2014 May;51(3):320-5. PubMed | Google Scholar
- Sarilita E, Setiawan AS, Mossey PA. Orofacial clefts in low- and middle-income countries: a scoping review of quality and quantity of research based on literature between 2010-2019. Orthod Craniofac Res. 2021 Aug;24(3):421-429. PubMed | Google Scholar
- Bentley AR, Callier SL, Rotimi CN. Evaluating the promise of inclusion of African ancestry populations in genomics. NPJ Genom Med. 2020;5:5. PubMed | Google Scholar
- Booth A, Clarke M, Dooley G, Ghersi D, Moher D, Petticrew M et al. The nuts and bolts of PROSPERO: an international prospective register of systematic reviews. Systematic reviews. 2012 Feb 9;1:2. PubMed | Google Scholar
- Moher D, Shamseer L, Clarke M, Ghersi D, Liberati A, Petticrew M et al. Preferred reporting items for systematic review and meta-analysis protocols (PRISMA-P) 2015 statement. Syst Rev. 2015 Jan 1;4(1):1. PubMed | Google Scholar
- Hupe M. EndNote X9. Journal of Electronic Resources in Medical Libraries. 2019;16(3-4):117-9. Google Scholar
- Butali A, Mossey PA, Adeyemo WL, Eshete MA, Gowans LJJ, Busch TD et al. Genomic analyses in African populations identify novel risk loci for cleft palate. Hum Mol Genet. 2019 Mar 15;28(6):1038-51. PubMed | Google Scholar
- Butali A, Mossey P, Tiffin N, Adeyemo W, Eshete M, Mumena C et al. Multidisciplinary approach to genomics research in Africa: the AfriCRAN model. Pan African Medical Journal. 2015 Jul 30;21:229. PubMed | Google Scholar
- Tishkoff SA, Reed FA, Friedlaender FR, Ehret C, Ranciaro A, Froment A et al. The genetic structure and history of Africans and African Americans. Science. 2009 May 22;324(5930):1035-44. PubMed | Google Scholar
- Grueber STaM. The economic impact and functional applications of human genetics and genomics. American Society of Human Genetics. 2021. Google Scholar